The highlights of Leroy Hood’s scientific career are like peaks in a mountain range spanning diverse fields, from molecular immunology and engineering, to genomics, to systems medicine. But Hood doesn’t think his trailblazing approach should be unusual, emphasizing that “one of the really key things about science is every 10 or 15 years, you really make a dramatic break and do something new… and you have to learn a lot before you can make fundamental contributions.”
One of Hood’s early contributions was in cracking the long-standing mystery of how the immune systems of humans and all vertebrates give rise to the vast diversity of antibodies that is critical for fighting myriad pathogens and foreign substances. During his PhD research at the California Institute of Technology (Caltech) in the mid-1960s, Hood and his advisor, William Dreyer, determined the amino acid sequences of components of antibody molecules and found that their sequences varied greatly between different antibodies. The finding helped advance their idea that each antibody is actually encoded by more than one gene, a big challenge to the existing dogma. Later, with his own research group at Caltech, Hood detailed the intricate process of how segments of antibody-encoding genes are rearranged, further creating antibody diversity. For his work in this area, Hood, along with Philip Leder and Susumu Tonegawa, won the Albert Lasker Basic Medical Research Award in 1987.
 Despite his groundbreaking discoveries in molecular immunology, Hood has not focused on this field for the last 20 years or so. Soon after starting his research group in 1970, he realized there were “striking technological limitations” that were holding scientists back from deeper understanding of the immune system. So, over the next two decades, Hood and his colleagues set out developing instruments, including the first automated DNA sequencer that allowed more rapid reads of genes. Hood’s intense focus on technology development as a faculty member at Caltech opened his eyes to what was the first paradigm shift of his career: “bringing engineering to biology.”
Despite his groundbreaking discoveries in molecular immunology, Hood has not focused on this field for the last 20 years or so. Soon after starting his research group in 1970, he realized there were “striking technological limitations” that were holding scientists back from deeper understanding of the immune system. So, over the next two decades, Hood and his colleagues set out developing instruments, including the first automated DNA sequencer that allowed more rapid reads of genes. Hood’s intense focus on technology development as a faculty member at Caltech opened his eyes to what was the first paradigm shift of his career: “bringing engineering to biology.”
Inspired by His Brother
Hood’s career path, traversing the worlds of biology, engineering, and other fields, seems like it was set at a young age. Hood spent much of his childhood hiking in the mountains in his native Montana, where his grandfather had a ranch. As a high school student, he participated in a summer geology camp at the ranch for university students and professors and won a Westinghouse Science Talent Search for his project mapping oil-producing rock formations. More than the accolade itself, Hood was struck by the diversity of the projects of the 39 other national winners he met in Washington, DC.
Throughout Hood’s earliest dabbles in science, and then his undergraduate degree at Caltech, he was drawn to human biology and medicine. He thinks this interest was engrained in him when he was 6 years old and had a brother born with Down syndrome. “What shocked me was when I asked several physicians why this occurred, they had not the faintest idea, and it was just an enormous mystery about why this fourth child could be so strikingly different… at that time, in 1944 or so, chromosomes hadn’t even been discovered, so we had very little idea,” Hood said. Hood attended a three-year medical degree program at Johns Hopkins Medical School to learn all he could about human biology. Although it was essentially an interlude between his undergrad and PhD studies at Caltech, medical school altered the path of Hood’s career in one major way: the microbiology course turned him on to immunology and steered his graduate research toward antibody diversity.
Around the time Hood was looking for PhD programs, William Dreyer was starting a lab at Caltech to focus on just what Hood wanted to study. Hood recalls that Dreyer told him early in grad school that his research to determine antibody characteristics “will be a simple Saturday afternoon project” because it was straightforward and there was not much competition. His mentor could not have been more wrong: within a year of embarking on the research, Hood says understanding antibody diversity became “one of the hottest areas in immunology,” and it stayed that way for the next decade.
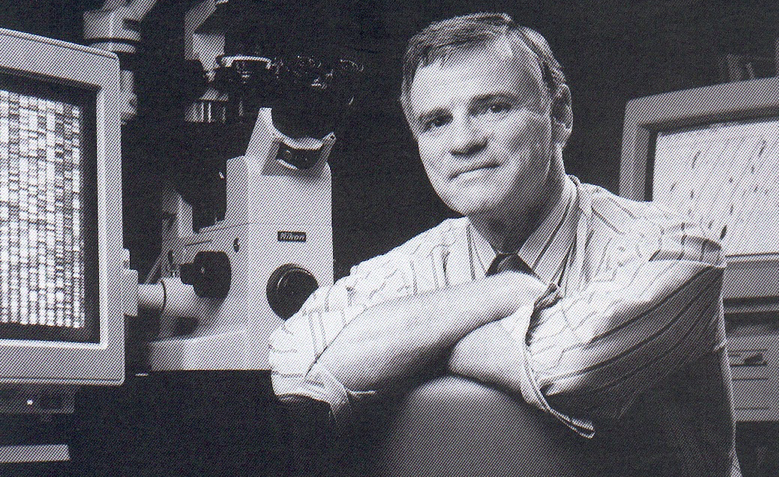

Hood with one of the early sequencers he helped develop
Designing New Technologies
During grad school, Hood got some other advice from Dreyer that ended up being a bit more on the money than his “Saturday afternoon project” prediction. While chatting about Dreyer’s own inventions, such as one of the first amino acid analyzers, Dreyer told Hood:
“If you want to practice biology, practice it at the leading edge, and if you want to transform a field, invent the technology that opens up new possibilities for generating data.”
When Hood returned from a stint at the Public Health Service at the US National Institutes of Health to Caltech for a faculty position, he had ample time to develop technology, but little support from the school to commercialize the instruments so that the rest of the scientific community could use them. As Hood explained, it was very different at Caltech back then, in the late 1970s, with no technology transfer office like there is today. Hood tried to sell his instruments to 19 companies but was rejected by all of them. Instead, he connected with a venture capitalist and started his own company, Applied Biosystems, in 1981. Its first product, the automated protein sequencer, took off, and Hood has since started 14 other companies, including Amgen and Arivale. Looking back, he notes his obstacles on the road to commercialization were actually blessings: “If you’ve got truly revolutionary ideas, you probably ought to start a company to execute them rather than try to sell them to an existing company,” which will probably not give them the attention they deserve.
Technology development led Hood to new scientific discoveries and business ventures, and it also drew him into entirely new fields of study. His automated DNA sequencer earned him an invitation in 1985 to the first meeting in Santa Cruz, California, to debate whether to launch the Human Genome Project. Hood and the other 11 attendees at the meeting were split on whether sequencing the entire human genome was a worthwhile endeavor. Around 90% of biologists back then were sharply opposed to the project for various reasons, including the belief that it would not yield useful information, with presumably only 2% of the genome containing actual genes, as Hood recalled recently. Of course, supporters of the project, including Hood, eventually won out, and Hood’s Caltech lab became one of dozens of centers in the United States, Europe, and Asia that together sequenced the human genome, a feat that was completed in 2003.
[Courtesy of the Institute of System Biology]
After participating in the Human Genome Project and seeing the first large-scale automated DNA sequencer that came out of it, Hood became increasingly convinced of the need for cross-disciplinary biology that encompasses chemistry, computer science, and other fields. When he was unable to persuade his biology colleagues at Caltech that the department should embrace this focus, Hood moved to the University of Washington School of Medicine in 1992 to establish the Department of Molecular Biotechnology, which he calls the first such cross-disciplinary program.
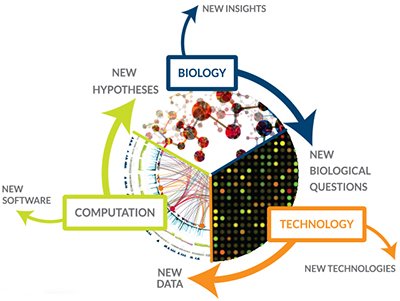
As Hood’s group at the University of Washington began collaborating with clinicians to apply genomics and proteomics to the study of prostate cancer and autoimmune disease, Hood became more and more interested in bringing medicine into the cross-disciplinary fold. At the same time, he believed that it was essential to study entire systems in biology — whether that be a cell, an organ, or a group of people — instead limiting one’s focus to individual genes or proteins. Hood coined a term for this approach, which is now a household name in biology departments: systems biology.
In 2000, Hood left the University of Washington to expand his systems biology focus and, together with colleagues Alan Aderem and Ruedi Aebersold, founded the Institute for Systems Biology (ISB), an independent nonprofit research institute in Seattle. As Hood reflected in a recent article about ISB, “[m]y colleagues and I began to create new technologies and strategic approaches to address [biological] complexity.” There are now hundreds of research institutes and centers worldwide taking a similar approach, Hood says.

Hood with a student in the lab
A Commitment to STEM Education
Just as Hood has been expanding the scope of his biological research, he and his wife Valerie Logan have been growing their efforts in science education. At ISB, they created the Logan Center for Education, which offers teacher training in STEM (science, technology, engineering, and mathematics) and provides resources for science faculty at community and technical colleges in Washington. Hood views education as one of academic scientists’ responsibilities: “We must train students to use inquiry-based analyses.”
Hood stepped down as president of ISB last year, but he still oversees a lab focusing on neurodegeneration and the science of wellness. In 2016, ISB began partnering with Washington-based Providence St. Joseph Health system, and Hood became their senior vice president and chief science officer to advance what he calls systems medicine. This approach essentially brings systems biology into the clinic and entails collecting a vast amount of data for each individual, including genome sequencing, to predict and prevent diseases.
Hood is attacking his current work with the same vigor he has had for more than half a century. “This is the most exciting time in my career, and I believe I still have a good 20 years ahead of me.”
By Carina Storrs
