Stephen J. Elledge
Brigham and Women’s Hospital

Evelyn M. Witkin
Rutgers University
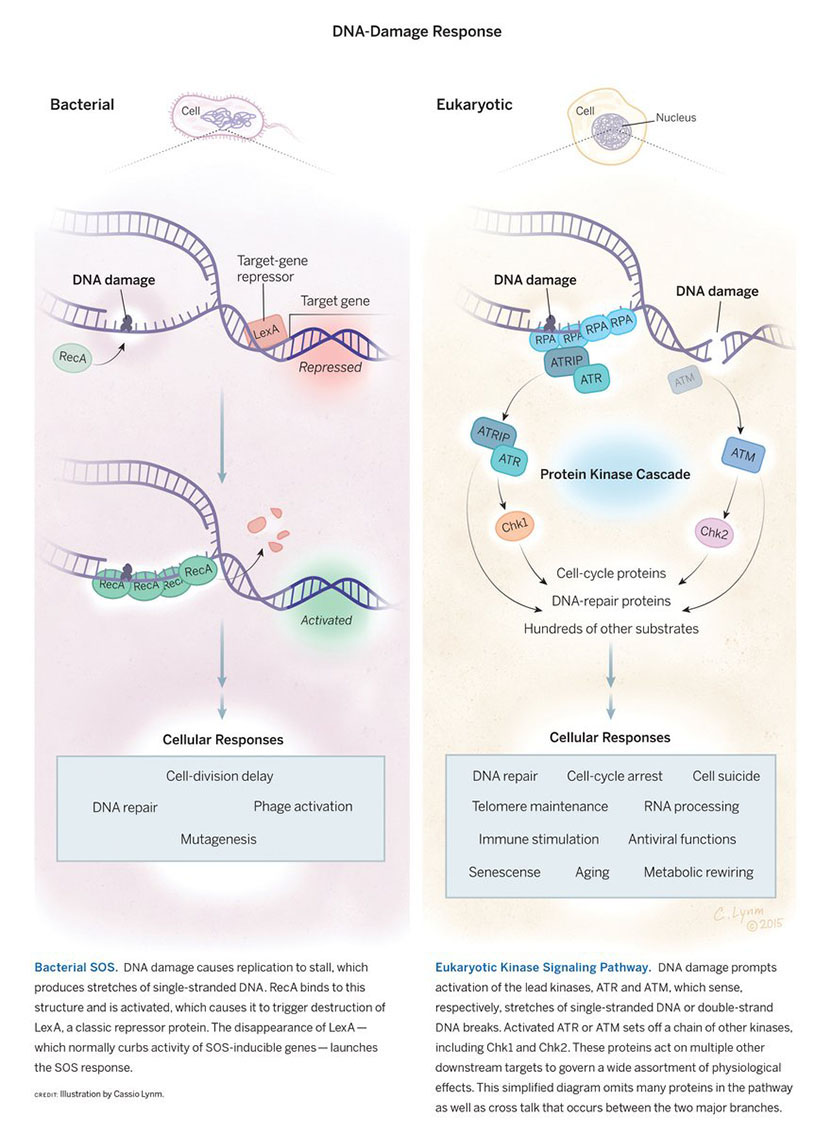
The 2015 Albert Lasker Basic Medical Research Award honors two scientists for their discoveries concerning the DNA-damage response, a mechanism that protects the genomes of all living organisms. Evelyn M. Witkin (Rutgers University) established its existence and basic features in bacteria, and Stephen J. Elledge (Brigham and Women’s Hospital) uncovered its molecular pathway in more complex organisms. The details of the two systems differ dramatically, yet they share an overarching principle. Both coordinate the activity of a large number of genes whose products shield creatures from potentially lethal harm.
Throughout their lives, cells withstand an onslaught of insults to their DNA. External agents such as chemicals and radiation bash it, and it also sustains abuse from within when normal physiological processes blunder. In humans, such events deliver tens of thousands of genetic wounds every day. The DNA-damage response detects not only DNA anomalies, but also other dangers, such as interruptions in the DNA-copying process. Living creatures then implement a multi-pronged strategy to ensure survival.
Award presentation by Titia de Lange
The oldest written texts are 5000 years old and are only known today because their Sumerian authors used clay tablets and kiln-baked their texts into permanence. The Etruscans hammered their best ideas onto gold plates, which also survived thousands of years. The progression to papyrus and paper often made written words short-lived and now the transition to electronic text has reduced that fleeting time to a New York minute.
What we write on our iPads is doomed to disappear fast, and baking our tablets in a kiln will not bring permanence. Of course the same changes that shortened the half-life of our texts came with enormous improvements in multiplication and distribution. But we may want to consider some mechanism to enhance the durability of what we write, even if most of it does not seem worth preserving.
Acceptance remarks
Acceptance remarks, 2014 Lasker Awards Ceremony
I got hooked on science in part because of the space race in the 1960s. There was a push to more broadly educate kids in science, and consequently I was exposed to the SRA reading laboratory in school. The advanced material was on science — fossils, planets, atoms and molecules. I loved atoms. The idea that matter was composed of smaller and smaller units with definable properties was astonishing. The fact that elements fell into the periodic table in a way that could predict their chemical properties is an amazingly elegant fact of nature that drew me in and continues to astound me even today. Initially, though, I avoided biology. But my senior year in college I learned about recombinant DNA — biology had gone molecular and that intersected with my interests in chemistry. The more I learned about DNA, the more I became seduced. In my mind, there is no more beautiful piece of art than the double helical structure of DNA. It is beauty and elegance personified. I wanted to be part of that revolution.
I was fortunate to be accepted to MIT without a biology background. I can remember when I first began doing research at MIT. I could not wait to get into the lab look at the results of my genetic experiments — did it work? I was a kid in a candy store and I just wanted to spend all of my time testing new ideas in the lab. It is an indescribable feeling of enlightenment when the right idea clicks into place to explain a phenomenon, an incredible rush. I am especially thrilled to share this award with Evelyn Witkin, whose pioneering work I learned of at MIT. While our careers barely overlapped, our interests were congruent. Can DNA sense its own integrity? What does it do with that information? We were both incredibly fortunate to work on such a rich and rewarding project.
There is nothing I would rather do than to be a scientist. It is a privilege to be able to pluck one of Mother Nature’s problems out of its firmament and examine how it works before putting it back, hopefully as a now shinier object. And to have something good come out of this work, to impact health and society, is more than I could have ever imagined. I am so grateful to have been able to play a role in science.
For me, the Lasker Award is a profound personal honor. I know that it recognizes not just the work of many creative individuals in my lab, but it is also a celebration of science itself. It is important for society to promote the culture of science and celebrate its values, the values of reasoning, openness, tolerance, and respect for evidence. That is why I so value the Lasker and its heritage. Mary Lasker saw far into the future and convinced this country of the importance of science and basic research, making a tremendous impact on society at large. By continuing to celebrate stellar science year after year, the Lasker Foundation is promoting the values of science.
I would like to end with a quote from Albert Einstein:
“One thing I have learned in a long life: That all our science, measured against reality, is primitive and childlike — and yet it is the most precious thing we have.”
Acceptance remarks, 2014 Lasker Awards Ceremony
I am deeply grateful to the Lasker Foundation and the Lasker Jury for the huge honor of this Award, which I am proud to share with Steven Elledge. I admit, though, that it feels slightly wicked to be recognized for work that was a labor of love and has always been its own reward.
John Tyndall, the 19th-century Irish physicist, in describing the mind of the scientist, wrote: “He lives a life of the senses, using his hands, eyes and ears in his experiments, but is constantly being carried beyond the margin of the senses. His mind must realize the subsensible world, and possess a pictorial power; if the picture so formed be correct, the phenomenon he is investigating is accounted for. Imagination with him does not sever itself from the world of fact; this is the storehouse from which all its pictures are drawn…”
I found that John Tyndall’s “pictorial power” always played a part in my scientific style. By 1961, after ten years of investigating the mechanism of ultraviolet light- (UV) induced mutagenesis in Escherichia coli, I had reached a conclusion. My experiments had told me that a UV-induced mutation occurs when an unrepaired UV photoproduct, most likely a pyrimidine dimer, prompts a replication error during the first DNA replication following UV exposure. I published those experiments and that conclusion, but not the vivid images that had formed in my mind, of just how that might happen.
In the 1960s, only one of E. coli’s five DNA copying enzymes, the DNA polymerases, was known, the one we now call Pol I. We also knew that this DNA polymerase was stopped cold by an unrepaired pyrimidine dimer in the DNA. That’s where imagination took over. I could almost ‘see’ another DNA polymerase, less fastidious than Pol I, knocking PoI I off the DNA and taking over as the copying enzyme, inserting any base at random, with a high probability of error, opposite the noncoding UV photoproduct.
In 1967, when I found that a mutation in the lexA gene eliminated UV mutability, I proposed that this gene encodes or controls just such a new DNA polymerase, responsible for UV mutagenesis by performing what we now call error-prone translesion synthesis.
In the early 1970s, Miroslav Radman and I pooled our separate observations and insights and recognized that UV damage induces a cluster of coordinately regulated, DNA damage-inducible activities, which came to be known as the SOS response. In time, dozens of E. coli genes were identified as SOS-inducible, including the umuC and umuD genes, essential for UV mutagenesis. It began to be possible to picture particular proteins carrying out translesion synthesis.
Harrison (Hatch) Echols, before his untimely death in 1983, was engaged in reconstructing SOS mutagenesis in vitro, with purified proteins. Hatch’s project has been carried on by Myron Goodman, Roger Woodgate, and coworkers at UCLA. They have identified the SOS-inducible error-prone DNA polymerase as Pol V, and have shown that it is composed of a particular arrangement of the UmuD and UmuC proteins. Six years ago, they showed that the “mutasome,” the active complex that performs SOS-induced translesion synthesis, is made up of Pol V+ RecA protein +ATP.
By this time, I was enjoying not just a visual image of SOS mutagenesis. It was more like watching an animated video with multiple components shown in such beautiful molecular detail that it was, for me, like seeing a long-imagined dream come true. I’m very glad that I lived to see it, and so sorry that Hatch didn’t.
Interview with Evelyn Witkin and Stephen Elledge
Video Credit: Flora Lichtman